| DNA2 |
|---|
|
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
|
|
| Identifiers |
|---|
| Aliases | DNA2, DNA2L, hDNA replication helicase/nuclease 2 |
|---|
| External IDs | OMIM: 601810; MGI: 2443732; HomoloGene: 6124; GeneCards: DNA2; OMA:DNA2 - orthologs |
|---|
| Gene location (Human) |
|---|
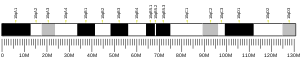
 | | Chr. | Chromosome 10 (human)[1] |
|---|
| | Band | 10q21.3 | Start | 68,414,064 bp[1] |
|---|
| End | 68,472,121 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 10 (mouse)[2] |
|---|
| | Band | 10|10 B4 | Start | 62,782,805 bp[2] |
|---|
| End | 62,809,964 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - secondary oocyte
- gonad
- testicle
- ventricular zone
- ganglionic eminence
- right lobe of liver
- Epithelium of choroid plexus
- rectum
- trabecular bone
- stromal cell of endometrium
|
| | Top expressed in | - fetal liver hematopoietic progenitor cell
- primitive streak
- abdominal wall
- otic placode
- epiblast
- otic vesicle
- tibiofemoral joint
- migratory enteric neural crest cell
- mandibular prominence
- medial ganglionic eminence
|
| | More reference expression data |
|
|---|
| BioGPS |  | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - DNA binding
- 4 iron, 4 sulfur cluster binding
- nucleotide binding
- helicase activity
- iron-sulfur cluster binding
- metal ion binding
- 5'-flap endonuclease activity
- DNA helicase activity
- ATPase activity
- protein binding
- catalytic activity
- 5'-3' DNA helicase activity
- nuclease activity
- endonuclease activity
- site-specific endodeoxyribonuclease activity, specific for altered base
- hydrolase activity
- ATP binding
- single-stranded DNA helicase activity
- RNA binding
| | Cellular component | - nucleoplasm
- gamma DNA polymerase complex
- mitochondrial nucleoid
- nucleus
- mitochondrion
- cytoplasm
| | Biological process | - mitotic telomere maintenance via semi-conservative replication
- mitochondrial DNA repair
- nucleic acid phosphodiester bond hydrolysis
- DNA double-strand break processing
- mitochondrial DNA replication
- cellular response to DNA damage stimulus
- DNA replication checkpoint signaling
- positive regulation of DNA replication
- metabolism
- base-excision repair
- DNA replication, removal of RNA primer
- DNA replication
- telomere maintenance
- DNA duplex unwinding
- DNA repair
- G-quadruplex DNA unwinding
- t-circle formation
- telomere maintenance via semi-conservative replication
- DNA replication, Okazaki fragment processing
- replication fork reversal
- regulation of signal transduction by p53 class mediator
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | | |
|---|
| RefSeq (protein) | | |
|---|
| Location (UCSC) | Chr 10: 68.41 – 68.47 Mb | Chr 10: 62.78 – 62.81 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|