| RECQL5 |
|---|
|
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
4BK0, 5LB8, 5LB5, 5LBA, 5LB3 |
|
|
| Identifiers |
|---|
| Aliases | RECQL5, RECQ5, RecQ like helicase 5 |
|---|
| External IDs | OMIM: 603781; MGI: 2156841; HomoloGene: 31232; GeneCards: RECQL5; OMA:RECQL5 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 17 (human)[1] |
|---|
| | Band | 17q25.1 | Start | 75,626,845 bp[1] |
|---|
| End | 75,667,189 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 11 (mouse)[2] |
|---|
| | Band | 11|11 E2 | Start | 115,783,421 bp[2] |
|---|
| End | 115,824,303 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - right lobe of thyroid gland
- right uterine tube
- left lobe of thyroid gland
- left testis
- anterior pituitary
- right testis
- sural nerve
- skin of abdomen
- body of stomach
- skin of leg
|
| | Top expressed in | - granulocyte
- zygote
- spermatid
- neural layer of retina
- secondary oocyte
- lip
- tail of embryo
- corneal stroma
- ventricular zone
- plantaris muscle
|
| | More reference expression data |
|
|---|
| BioGPS | 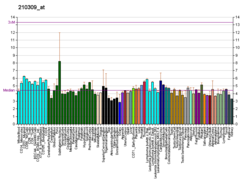
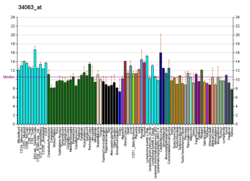
 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - RNA polymerase II complex binding
- nucleotide binding
- helicase activity
- DNA helicase activity
- nucleic acid binding
- hydrolase activity
- ATP binding
- DNA binding
- 3'-5' DNA helicase activity
- four-way junction helicase activity
- protein binding
- identical protein binding
| | Cellular component | - RNA polymerase II, holoenzyme
- nucleoplasm
- nucleus
- chromosome
- cytoplasm
- cytosol
| | Biological process | - DNA recombination
- DNA metabolic process
- negative regulation of transcription elongation from RNA polymerase II promoter
- cellular response to DNA damage stimulus
- cell division
- chromosome separation
- DNA replication
- cell cycle
- DNA repair
- DNA duplex unwinding
- double-strand break repair via homologous recombination
- cellular response to camptothecin
- replication-born double-strand break repair via sister chromatid exchange
- negative regulation of double-strand break repair via homologous recombination
- mitotic cell cycle
- mitotic sister chromatid segregation
- response to X-ray
- negative regulation of mitotic cell cycle
- DNA unwinding involved in DNA replication
- metabolism
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_001003715
NM_001003716
NM_004259 |
| |
|---|
| RefSeq (protein) | |
|---|
NP_001003715
NP_001003716
NP_004250 |
| |
|---|
| Location (UCSC) | Chr 17: 75.63 – 75.67 Mb | Chr 11: 115.78 – 115.82 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|


















